What to know about COVID-19 (SARS-CoV-2) Omicron Variant Sub-Lineages?

Understanding the evolution of Omicron and its sub-lineages can provide insight into the risk of another “Omicron”-type variant. In this paper we examine in every aspect of what to know about COVID-19 (SARS-CoV-2) Omicron sub-variants.
COVID-19 Omicron BA.2 & BA.3 Sub-variants
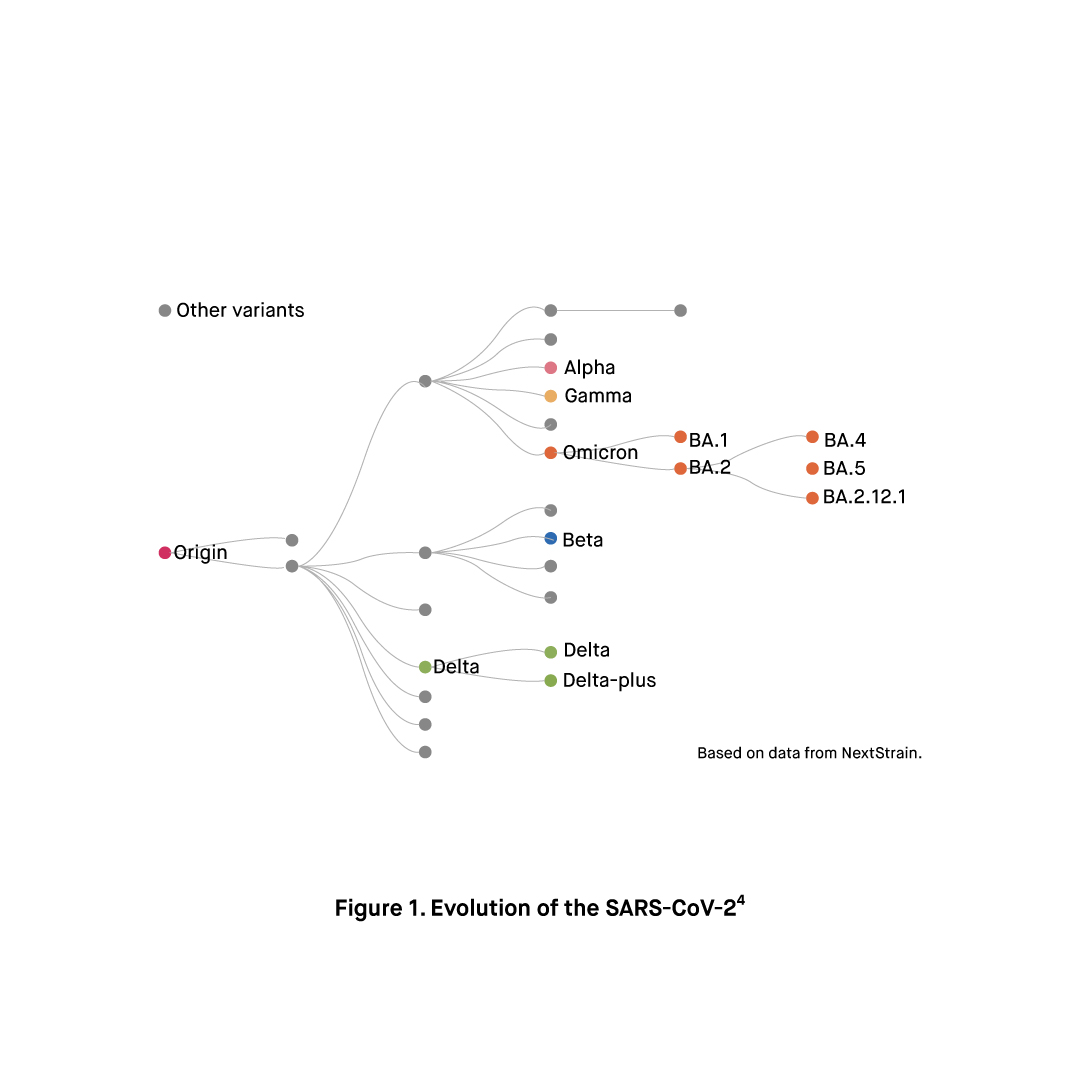
The BA.3 sub-lineage has been first detected in southern Africa on November 18, 2021, and as of 18 April 2022 2,875 BA.3 sequences in total have been reported from at least 64 countries. The majority of the BA.3 sequences have been reported from the United Kingdom (648), Poland (452), India (427), Germany (373), Israel (180), South Africa (166), and the United States (157).
The data on the infectiousness, disease severity, and immune evasion of the BA.3 sub-lineage is scarce. On one hand, in a pre-print in-silico study concerning spike mutation profiles, Kumar et al. argued that the BA.3 and BA.2 sub-lineages have slightly higher binding affinity to the human angiotensin converting enzyme-2 (hACE-2) receptor than wild-type, BA.1, and BA.1.1 lineages, which might indicate increased tropism and infectivity. In the same study, Kumar et al. have also estimated that the spike protein of the BA.3 sub-lineage would have slightly more hydrophobic residues than other Omicron sub-lineages, which may result in higher pathogenicity1. On the other hand, Desingu et al. has suggested that the absence of certain mutations found in the spike proteins of BA.1 and BA.2, might explain the relatively lower number of cases infected with the BA.3 sub-lineage2. As of now, the data remains insufficient to assess the infectivity and disease severity of the BA.3 sub-lineage.
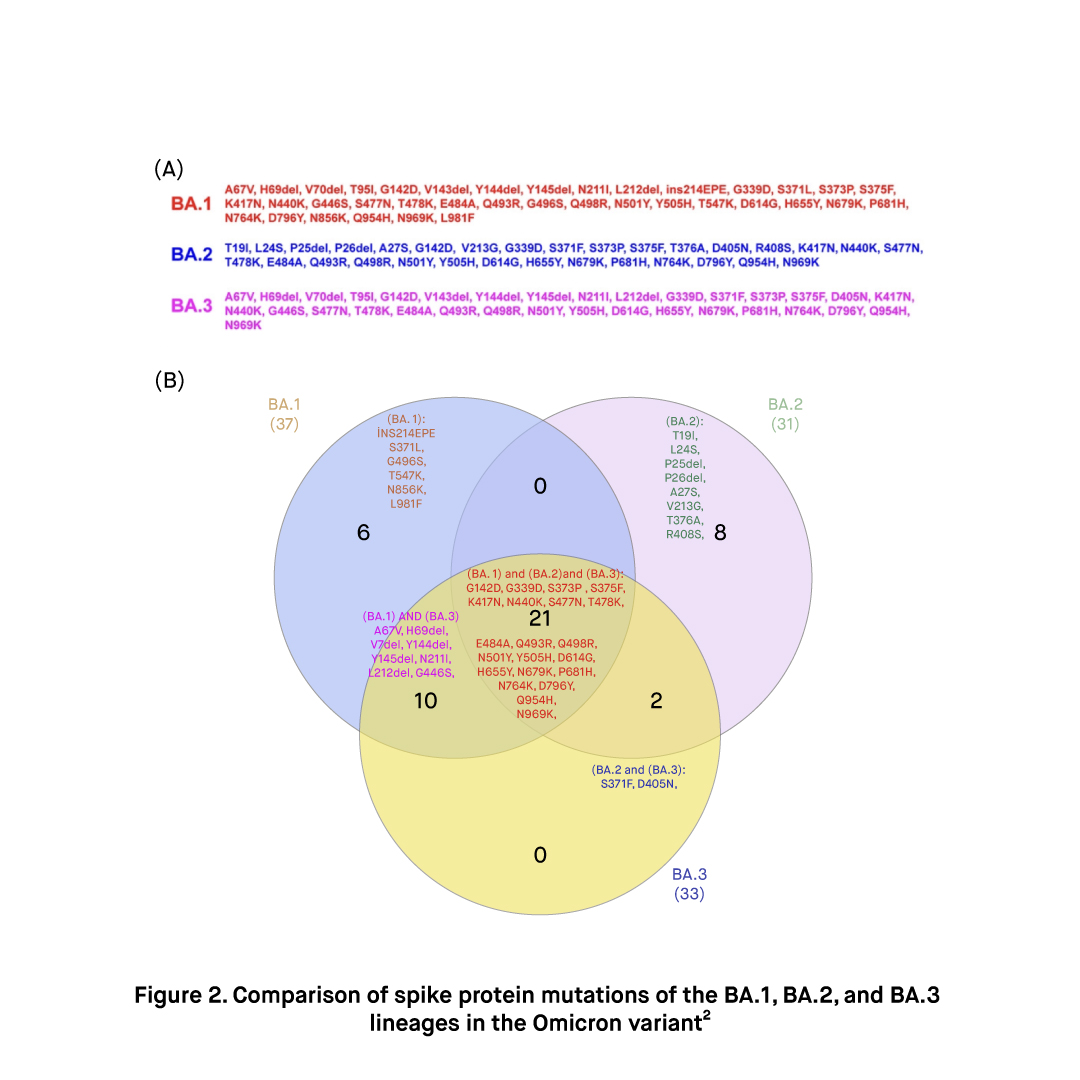
Concerning natural immunity and vaccine-induced immunity escape, limited in-vitro evidence indicates that the BA.3 variant may be more efficient relative to BA.1 and BA.2. For instance, utilizing neutralization assays and a pseudo-virus developed to express the spike proteins of these sub-lineages, it has been reported that the ability of the BA.3 to escape both vaccine-induced immunity offered by Pfizer-BioNTech Comirnaty COVID-19 vaccine and natural immunity may be higher than that of B.1 and BA.2, but comparable to that of BA.1. Another study by Kurhade et al. has reported that the BA.3 variant has demonstrated an increased ability of evading vaccine-elicited immunity provided by Pfizer-BioNTech Comirnaty COVID-19 vaccine compared to both BA.1 and BA.23.
While more data is required to evaluate the exact performance of our current diagnostic tests against the BA.3 sub-lineage, considering the mutations shared between the Omicron sub-lineages and reported cases, it is highly expected that both molecular tests such as PCR and antigen tests would have similar performances in the detection of the main Omicron sub-lineages.
COVID-19 Omicron BA.4 and BA.5 Sub-variants
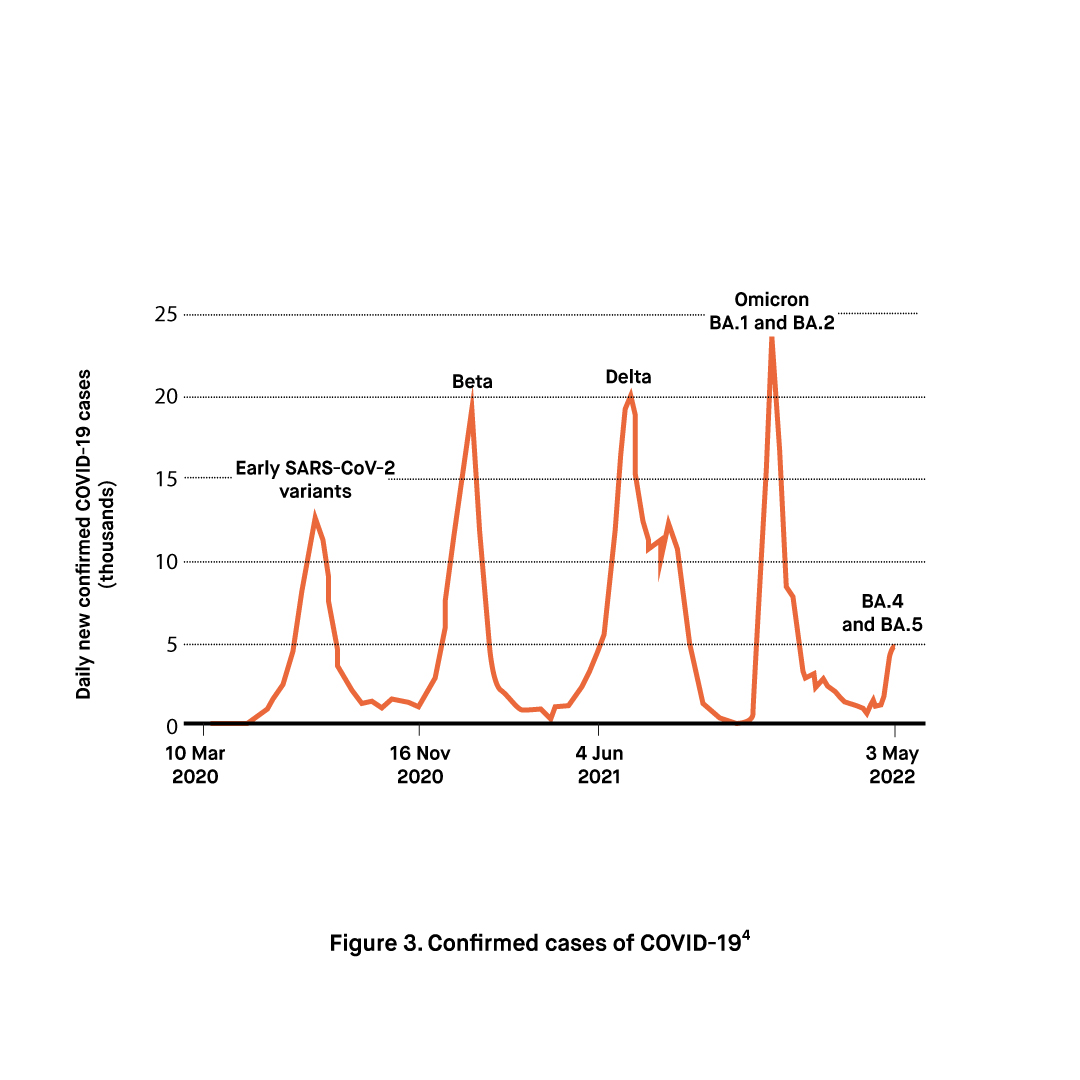
The World Health Organization has started to monitor Omicron sub-lineages BA.4 and BA.5 due to mutations L452R and F486V in their spike proteins which are linked to potential immune escape. Another noteworthy mutation is the 69/70 deletion shared by BA.4 and BA.5 lineages which function as a proxy indicator in the detection of the variant by enabling the S-gene target failure. Thus, unlike the BA.2 sub-lineage which does not contain the 69/70 deletion, the BA.4 and BA.5 variants can be identified without gene sequencing.
Classified in the United Kingdom on April 2, 2022, the BA.4 sub-lineage shares all mutations and deletions with BA.2 while also containing the 69/70 deletion, and L452R, F486V, Q493 (wild type) mutations in its spike protein, along with mutations NSP4: L438F, ORF 7b: L11F, N: P151S, and ORF 6: D61 (wild type) outside of its spike protein. The BA.5 sub-lineage shares the mutations and deletions with its sibling except the mutations M: D3N, ORF 7b: L11 (wild type), and N: P151 (wild type), along with synonymous SNPs: A27038G, C27889T outside of its spike protein.
As of 26 April 2022, 222 BA.4 sequences in total have been reported in South Africa, Denmark, United Kingdom, Belgium, United States, Austria, Australia, Botswana, France, Switzerland. The BA.5 sub-lineage has been less prevalent than its sibling with 99 sequences in total reported from South Africa, Germany, Portugal, United Kingdom, United States, France, Denmark, Hong Kong, and Spain. Israel has also reported three BA.4 and two BA.5 sequences on 25 April 2022.
The assessment of the BA.4 and BA.5 sub-lineages regarding viral transmission, disease severity, and immunity evasion is ongoing. As of now, the data remains insufficient to draw precise and reliable conclusions. However, according to the United Kingdom Health Security Agency (UKHSA), the geographic spread of the BA.4 sub-lineage may be an indicator of its ability of transmission. While the mutations L452R and F486V shared by the BA.4 and BA.5 sub-lineages are associated with potential immune escape, there is currently insufficient information to determine if or how infections with BA.4 or BA.5 might differ from infections with the BA.2 sub-lineage both in terms of immunity escape and disease severity.
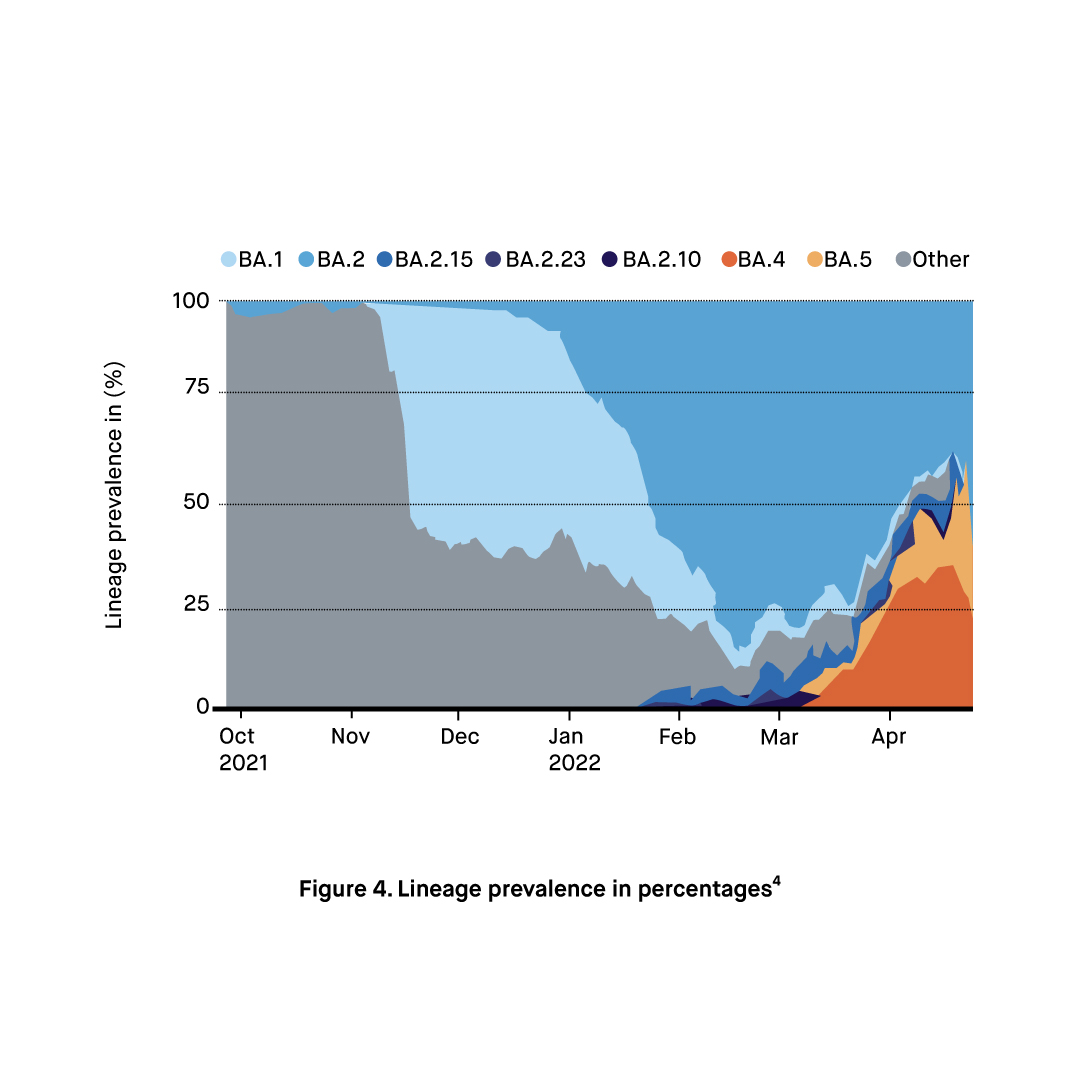
COVID-19 Omicron XE Sub-variants
First detected in the United Kingdom on January 19, 2022, the XE subvariant shares most mutations with the BA.2 sub-lineage while containing mutations from the BA.1 sub-lineage for nonstructural proteins 1-6, along with three other mutations differing from both the BA.1 and the BA.2. As of April 18, 2022, 1365 cases infected with the XE sub-lineage have been reported worldwide. The majority of these were reported from the UK with 1322 sequences. The rest was reported from the US (15), Israel (7), Germany (5), Denmark (4), Switzerland (3), Ireland (3), Australia (1), Brazil (1), Belgium (1), Sweden (1), India (1), and France (1). More recently, the media have also reported a number of cases infected with the XE sub-lineage in Canada, South Korea, Thailand, Northern Ireland, and New South Wales.
Preliminary data from UK-based community testing indicates that XE had a higher growth rate relative to the BA.2 subvariant. In fact, through logistical regression, the median growth rate of the XE sub-lineage have been found to be 12.6% higher within the period between January 15 and March 30, 2022. Within the last three weeks prior to 30 March, it was 20.9% higher than the median growth rate of the BA.2 sub-lineage. However, as variable growth rates have been observed among different areas within UK, further data is required to assess the growth rate of the XE sub-lineage. There is also no sufficient information to determine how and whether the XE sub-lineage differs from the BA.2 variant regarding disease severity and immunity escape.
Can COVID-19 Omicron Sub-Variants Be Detectable by Rapid Antigen Test Kits?
Antigens are proteins or protein fragments that act as a target for a specific antibody in a reactive test strip. Spike proteins and nucleocapsid proteins are examples of antigens that we use. The spike protein region is home to the majority of the mutations in the Omicron sub-variants. For rapid antigen tests using SARS-CoV-2 spike protein as a target, this can reduce sensitivity or result in non-functional tests. In terms of sensitivity and functionality, those that use nucleocapsid protein as a target, such as our RapidFor™ SARS-CoV-2 Rapid Antigen Test Kit, are less likely to be influenced by mutations.
References
- Kumar, S., Karuppanan, K., & Subramaniam, G. (2022). Omicron (BA.1) and Sub-Variants (BA.1, BA.2 and BA.3) of SARS-CoV-2 Spike Infectivity and Pathogenicity: A Comparative Sequence and Structural-based Computational Assessment. bioRxiv. https://doi.org/10.1101/2022.02.11.480029
- Desingu, P. A., Nagarajan, K., & Dhama, K. (2022). Emergence of Omicron third lineage BA.3 and its importance. Journal of Medical Virology, 94(5), 1808–1810. https://doi.org/10.1002/jmv.27601
- Kurhade, C., Zou, J., Xia, H., Cai, H., Yang, Q., Cutler, M., Cooper, D., Muik, A., Jansen, K. U., Xie, X., Swanson, K. A., & Shi, P. Y. (2022). Neutralization of Omicron BA.1, BA.2, and BA.3 SARS-CoV-2 by 3 doses of BNT162b2 vaccine. bioRxiv. https://doi.org/10.1101/2022.03.24.485633
- Callaway, E. (2022). Are COVID surges becoming more predictable? New Omicron variants offer a hint. Nature, 605(7909), 204–206. https://doi.org/10.1038/d41586-022-01240-x

