In a significant development, the United Kingdom has reported its first human case of Influenza A(H1N2)v, a variant of the swine flu virus. This discovery has triggered health officials to initiate thorough investigations and preventive measures to contain the potential spread of the virus.

What is Influenza A(H1N2)v?
Influenza A(H1) viruses are prevalent in swine populations globally, with subtypes like H1N1, H1N2, and H3N2 occasionally infecting humans. The newly detected A(H1N2)v variant, though similar to viruses found in UK pigs, presents a distinct clade (1b.1.1). This marks the first instance of such a virus in humans in the UK.

Symptoms and Epidemiology
Individuals infected with Influenza A(H1N2)v may experience respiratory symptoms akin to seasonal influenza, including fever, cough, and body aches. The individual in the UK case exhibited mild symptoms and has fully recovered. The epidemiology of this variant virus is under intense scrutiny, given its sporadic occurrence globally, with only 50 reported human cases since 2005.
Detection and Surveillance
The case was identified through routine national flu surveillance by the UK Health Security Agency (UKHSA) and the Royal College of General Practitioners (RCGP). The individual, who experienced mild respiratory symptoms, tested positive for Influenza A(H1N2)v using polymerase chain reaction (PCR) testing and genome sequencing.
Response and Investigation
UKHSA, in collaboration with partners, is actively working to assess the risk to human health and trace close contacts of the infected individual. The infected person, who has fully recovered, had no known contact with pigs, raising questions about the source of the infection. Investigations are underway to ascertain how the individual acquired the virus.
Health Measures and Advice
Close contacts are being monitored, tested as necessary, and provided with appropriate care. Individuals exhibiting respiratory symptoms are advised to follow existing guidelines, avoiding contact with others, especially vulnerable populations.
Expert Insights
UKHSA, highlights the role of routine flu surveillance and genome sequencing in detecting the virus. Investigations aim to understand the infection’s origin and assess the potential for further cases.

Chief Veterinary Officer, emphasizes the importance of high standards in animal health, welfare, and biosecurity. Collaboration between animal and human surveillance systems is crucial to protecting public health.
Global Significance
With only 50 cases of Influenza A(H1N2)v reported worldwide since 2005, the UK case raises awareness about the potential for transmission from animals to humans and the need for vigilant monitoring.

Background on Influenza Viruses
Influenza viruses, known for their ability to cause seasonal epidemics and occasional pandemics, belong to the family Orthomyxoviridae. These viruses are classified into types A, B, C, and D based on their core proteins. Influenza A viruses are the most diverse and are further categorized into subtypes based on their surface proteins, hemagglutinin (H) and neuraminidase (N).

Influenza A(H1) Viruses in Swine Populations:
Influenza A(H1) viruses are enzootic in swine populations worldwide. Enzootic refers to the continuous, low-level presence of the virus within the swine population. When an influenza virus that typically circulates in swine is identified in a human, it is termed a ‘variant influenza virus.
Swine Influenza A Virus Subtypes:
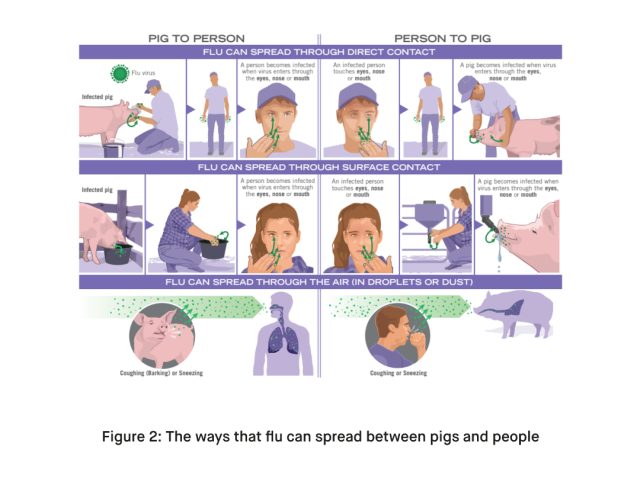
Swine influenza A viruses primarily include subtypes H1N1, H1N2, and H3N2. These subtypes circulate among pig populations and have the potential to infect humans. Human infections often occur following direct or indirect exposure to pigs or environments contaminated with swine influenza viruses.

Influenza A(H1N2)v: A Variant Strain:
In the context of the recent detection in the UK, Influenza A(H1N2)v is identified as a variant strain. This variant is similar to viruses currently circulating in pigs in the region. The designation ‘v’ denotes its origin in swine, emphasizing the zoonotic potential, where the virus can jump from animals to humans.
Genetic Clades and Global Context:
The genetic diversity of influenza viruses is further classified into clades. The infection detected in the UK is described as a distinct clade (1b.1.1), distinguishing it from recent human cases of Influenza A(H1N2) worldwide. This emphasizes the importance of genetic surveillance to track the evolution and spread of influenza viruses.
Historical Perspective: 2009 H1N1 Pandemic:
The 2009 influenza pandemic was caused by an influenza A virus (H1N1pdm09), commonly referred to as ‘swine flu.’ This virus contained genetic material from viruses circulating in pigs, birds, and humans in the 1990s and 2000s. While H1N1pdm09 now circulates seasonally in humans, it is distinct from the viruses currently circulating in pigs.
Understanding the background of influenza viruses, their prevalence in swine populations, and the occasional transmission to humans is crucial for surveillance, early detection, and effective management of emerging variants. Vigilant monitoring and collaboration between human and animal health systems remain integral in preventing and controlling potential outbreaks.

Which Diagnostic Test Kits use for the Influenza A virus?
Currently, there is no specificied commercially available diagnostic test kit for new variant Influenza A(H1N2)v strain. However, there is several commercialized rapid diagnostics test kits for detecting Influenza virus including Influenza A (H1N1), the dominant subtype of swine flu virus which is also called seasonal influenza.
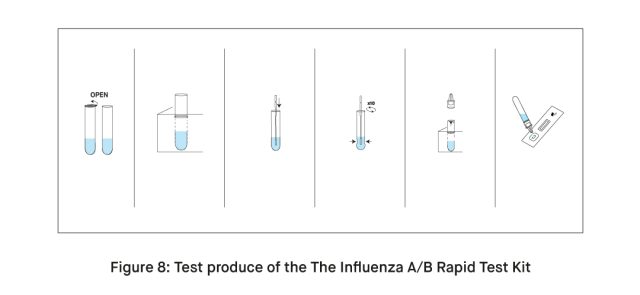
The Influenza A/B Rapid Test Kit offers a comprehensive solution by enabling the identification of both Influenza A and B viral nucleoprotein antigens in human nasopharyngeal and nasal swab samples. This 2-in-1 capability streamlines the diagnostic process for influenza infections. The utilization of nasopharyngeal swabs enhances the accuracy of sample collection. The test is designed for ease of use, ensuring a straightforward procedure for healthcare professionals. One of its notable advantages is the rapid turnaround time, providing results within 15 minutes of sample collection.
This expeditious reporting is crucial for prompt decision-making in clinical settings. Furthermore, the methodology employed by the test kit is not only efficient but also safe, contributing to its reliability in influenza diagnosis. Overall, the incorporation of these features positions the test kit as a valuable tool in the rapid and accurate detection of both Influenza A and B viruses.
REFERENCES
Harrington, W.N., Kackos, C.M. & Webby, R.J. The evolution and future of influenza pandemic preparedness. Exp Mol Med 53, 737–749 (2021). https://doi.org/10.1038/s12276-021-00603-0
Centers for Disease Control and Prevention. (2019, April 16). What CDC does about novel flu: Outbreak investigations. Centers for Disease Control and Prevention. https://www.cdc.gov/flu/outbreak-investigations.html
Centers for Disease Control and Prevention. (2023, March 30). Types of influenza viruses. Centers for Disease Control and Prevention. https://www.cdc.gov/flu/about/viruses/types.htm
Agency, U. H. S. (2023, November 27). UKHSA detects human case of influenza A(H1N2)V. GOV.UK. https://www.gov.uk/government/news/ukhsa-detects-human-case-of-influenza-ah1n2v


